Se170063
PAA_Selenium/Soil_Experiments#Selenium_Sample_Analysis
Sample Description
The sample was placed in an aluminum cylinder that was to be irradiated. The target components consisted of a nickel foil on the front of the cylinder with 2 pure selenium pellets under the foil, but still outside the cylinder. Inside the target there was burnt sagebrush ash, which was burned with a blowtorch, and selenium. Below are the masses of the components
Nickel Foil: 0.2783g
Outer Se Pellets: 0.0971g
Sage Ash: 0.5111g
Inner Se Pellets: 0.0523g
Energy
The Calibration for Detector A was done on the morning of 5/23/17 with the MPA software using the thorium rods (as the calibration was fairly close already) and the correction values were found to be Det A Intercept = -12.208800 slope =1.021270
Efficiency
Nickel Information
Activity and Half Life
Se170063 Activity And Half Life
Alternative Method
Se170063 Activity and HL Alternate
Corrected Alternative method
The files used for this analysis are in the directory /data/IAC/Se/May2017/5_25_17/Se_Activity_SysOffset_Mix
To begin, I have corrected the mixture to have a factor of 0.62, which is the mystery factor throwing of all of these analyses. The histogram is also weighted by the mass. The weight added to the histogram is
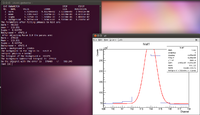
So the true number of counts has indeed been weighted here. Now I want to try to test every different method that was suggested. So first I am going to weight the mixture by the mystery factor of 0.62, and leave my Gaussian fits as wide as they were previously. The gaussians will probably be made more compact if the mystery factor does not alleviate the problem. The first step is to find the number of counts within the window of interest. Below is the process I used to determine the number of counts and the error associated with it. First begin by plotting the histogram using the ROOT program Eff.C, which is shown below.
Now take the integral given in the stats box and subtract the background to get the number of counts. Here the number of counts would be
Now I can convert this error into an activity by dividing by the time, which is 300 seconds in this case. After that take the natural log of the quotient.
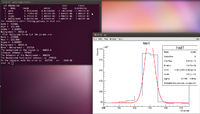
Now to find the error we can notice that the standard deviation here is 0.6238. The procedure is to expand the window by one or two standard deviations and find the difference in the number of counts in the original window. Due to the binning here I have decided to expand the window by 1 channel on each side, which is roughly 2 standard deviations. A picture is given below.
Now we do a similar method to find the number of counts within this window.
Now take the difference between the original window's number of counts and the expanded window's number of counts.
Now to propagate this error we must divide this number by the original number of counts.
This method was repeated for the next set of runs that make the data file. The numbers are in a table below.
| 0 <t< 300 sec | 730 < t < 1020 sec | 1480 < t < 1775 | 2250 < t < 2550 sec | 3050 < t < 3300 sec | 3775 < t < 4050 sec | 4480 < t < 4770 sec | |
| Original Window Counts | 880200 | 716200 | 617100 | 545800 | 394100 | 362000 | 346300 |
| Original Window Background | 47672.4 | 33451 | 26907.5 | 23708.2 | 17990.1 | 17192.7 | 16024.6 |
| Original Window Difference | 832527.6 | 682749 | 590192.5 | 522091.8 | 376109.9 | 344807.3 | 330275.4 |
| Expanded Window Counts | 970900 | 782600 | 671300 | 593100 | 426700 | 395400 | 378100 |
| Expanded Window Background | 46813.8 | 33363.3 | 26902.5 | 23647.6 | 17335.3 | 17008.9 | 15964.4 |
| Expanded Window Difference | 924086.2 | 735786.2 | 644397.5 | 569452.4 | 409364.7 | 378391.1 | 362135.6 |
| Error in counts | 91558,6 | 53037.2 | 97305.7 | 47360.6 | 15264.7 | 33583.8 | 31860.2 |
| .dat file entry | 7.928439179 +/- 0.1099766542 | 7.811833814 +/- 0.0776818421 | 7.573569236 +/- 0.1778599131 | 7.461816239 +/- 0.0907131658 | 7.316175749 +/- 0.040585749 | 7.133969892 +/- 0.0887542017 | 7.037801208 +/- 0.964655557 |
Runlist
Table with dates and filename and locations on daq1